2.4 Custom librariesYou can use a custom library to connect specific pairs of fragments. A custom library could include
Custom libraries are limited to linear spacers (no fuse-rings), and do not include at present group files. They are also stored differently than the default spacer library. Namely, the spacer coordinate files and the keyfile must reside in the same directory. You define the spacers for a custom library and you edit it with nfe as you would do for the default library. Take care to save the keyfile and the coordinates in a directory that can be accessed during a NEWLEAD session. Connecting fragments with custom librariesTo use one or more custom library, define the fragments and the preferences for NEWLEAD as usual. Decide then which custom library is to be used for a given pair of fragments. You can still use the default library for one or more pairs of fragments, when using custom libraries. stand alone use identify the sequence of each fragment in the input file for NEWLEAD. Add to the command line the sequence of pairs to be connected using the option -f, and the corresponding custom libraries using the keyword -c. The custom libraries are identified by their keyfiles. The character "&" identifies the default library, and the character "=" specifies that the previous library must be used for the next pair as well. For example
means that the library "my_spacers.lib" is used to connect both fragment 1 to 2 and 2 to 3.
means that the standard library is used to connect fragment 1 to 3, and the custom library "alkane3.lib" is used to connect fragment 1 to 2.
means that the custom library "lab2.lib" is used for the first two pairs as given, and "lab1.lib" is used for the last pair. You can also omit the keyword -f, but specify the keyword -c, in which case NEWLEAD will decide which pair to connect in each pass according to its internal rules, but will use the specified libraries for the connections. In such cases, it is meaningful to use only one library, e.g.
uses a custom library in all passes (extra "=" characters are ignored). |
||
|
Omitting the -c keyword and using the -f keyword forces the connection of fragments in a given order, with the default library, e.g.
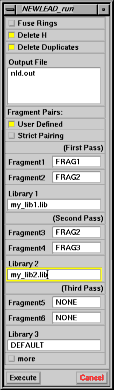
forces the connection of fragments 1 and 3 first, and fragments 1 and 2 last. This option, not related to custom libraries, can be useful to speed up a run by attempting first the connection of the fragment pair which is most difficult to connect. Accelrys interface Activate the NEWLEAD_run command, and select the "User Defined" button for fragment connections. The window expand as shown in Fig. 6. You see the following buttons and fields
To fill in the fragment fields, pick molecules on the screen, or type the corresponding names. Leave the word "NONE" if you want NEWLEAD to decide the connections for a given pass. Type the full pathnames of the library keyfiles in the library fields. Leave the word DEFAULT for the default library. You can also use the character "=" to use a second time the library in the previous field. |
|
| |
|